About
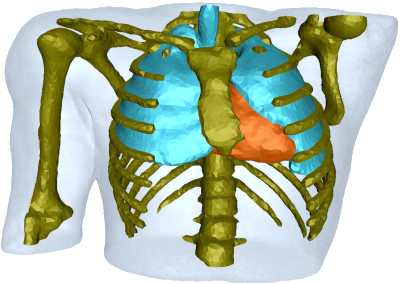 Workshop on Mesh Processing in Medical Image Analysis
Workshop on Mesh Processing in Medical Image AnalysisIn conjunction with MICCAI 2011.
Westin Harbour Castle, Toronto, Canada
September 18th, 2011
Scope
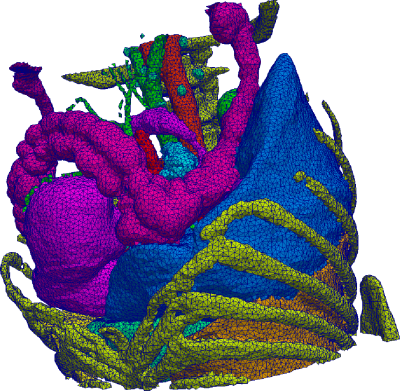 Many strategies for medical image analysis have been built on an
image analysis pipeline that starts with acquired image data,
performs filtering and processing, constructs geometric models of
important surfaces and structures, performs simulation, and finally
provides quantitative and visual analysis of the data. Within this
pipeline, geometry and shape are commonly represented as a mesh, or a
discretization of some domain into simpler computational elements such as
quads or triangles (representative of surface pieces) or tetrahedra and
hexahedra (representative of volumetric elements). This
image-to-mesh (I2M) step converts volumetric images into formats
that are more suitable for solving finite-element simulations, analyzing
critical structures, and performing boundary surface visualization tasks.
Current research in computational geometry, graphics hardware, and
computer graphics has produced methods to represent, extract, refine,
visualize and analyze both critical surfaces embedded in the 3D volumes,
such as interfaces between tissues, as well as volumetric regions, such as
organs.
Many strategies for medical image analysis have been built on an
image analysis pipeline that starts with acquired image data,
performs filtering and processing, constructs geometric models of
important surfaces and structures, performs simulation, and finally
provides quantitative and visual analysis of the data. Within this
pipeline, geometry and shape are commonly represented as a mesh, or a
discretization of some domain into simpler computational elements such as
quads or triangles (representative of surface pieces) or tetrahedra and
hexahedra (representative of volumetric elements). This
image-to-mesh (I2M) step converts volumetric images into formats
that are more suitable for solving finite-element simulations, analyzing
critical structures, and performing boundary surface visualization tasks.
Current research in computational geometry, graphics hardware, and
computer graphics has produced methods to represent, extract, refine,
visualize and analyze both critical surfaces embedded in the 3D volumes,
such as interfaces between tissues, as well as volumetric regions, such as
organs.
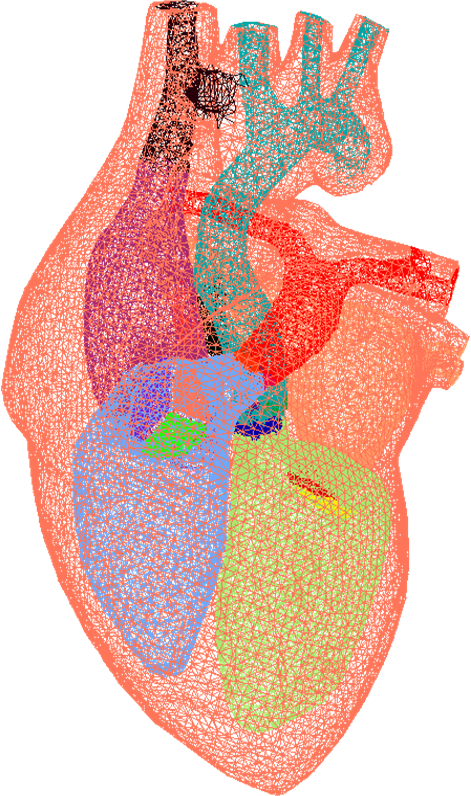 The workshop investigates the role meshes have with medical image
analysis and is broadly based on three overlapping themes:
The workshop investigates the role meshes have with medical image
analysis and is broadly based on three overlapping themes:
- Mesh processing,
- The I2M pipeline,
- Surface analysis and extraction.
Best paper prize
The MeshMed 2011 best paper prize was awarded to Katarzyna Wełnicka, Jakob Andreas Bærentzen, Henrik Aanæs, and Rasmus Larsen for the paper Example based style classiffication.Registration
Registration for the workshop is handled trough the main MICCAI conference. The deadline for early-bird registration is August 1. On site registration is in the Frontenac Foyer. The registration opening times:- Sun Sept 18: 7:00-17:00
- Mon Sept 19: 7:00-17:00
- Tue Sept 20: 7:00-17:00
- Wed Sept 21: 8:00-17:00
- Thu Sept 22: 8:00-11:00
Organisers

- Rasmus R. Paulsen (Technical University of Denmark)
- Joshua A. Levine (SCI Institute, University of Utah)
- Christian Barillot (IRISA/CNRS/INRIA Rennes, France)
- Nikos P. Chrisochoides (Old Dominion University)
- Hervé Delingette (Asclepios, INRIA Sophia-Antipolis, France)
- Ross T. Whitaker (SCI Institute, University of Utah)
- Yongjie Zhang (Carnegie Mellon University )
Sponsors
The workshop is sponsored by the NIH/NCRR Center for Integrative Biomedical Computing at the Scientific Computing and Imaging Institute, University of Utah and the Department of Mathematical Modeling at the Technical University of Denmark.Contact
|
Rasmus R. Paulsen Associate Professor Informatics and Mathematical Modelling Technical University of Denmark Richard Petersens Plads Building 321, office 219 DK-2800 Kgs. Lyngby Denmark Phone: +45 4525 3423 Fax: +45 4588 1397 www: http://www.imm.dtu.dk/~rrp Email: rrp at imm dot dtu dot dk |
Joshua A. Levine Postdoctoral Research Associate Scientific Computing and Imaging Institute University of Utah 72 S Central Campus Drive WEB 2815 Salt Lake City, UT 84112 USA Phone: +1 801-585-1867 Fax: +1 801 585 6813 www: http://www.sci.utah.edu/~jlevine Email: jlevine at sci dot utah dot edu |

