
The lectures are given in building 321, room H033, on Thursdays from 1 pm – 2.30 pm (approximately). The computer exercises take place immediately after the lectures in the same room; a teaching assistant will be present. The computer exercises are on the topic of the current week’s lecture. The exercises should be carried out in groups of 2-3.
Lecture notes and exercise texts and materials are available through CampusNet.
For each subject considered in the course an exercise is carried out and an exercise report must be prepared and handed in. The course examination consists of an oral examination in the course topics based on the course theory and exercise reports. The oral examination will be held on Tuesday, 10 December 2019.
This is a 5 ECTS point course corresponding to a work load of 8-10 hours work per week and most of this time is used to implement the image analysis models in the exercises. Some of the algorithms are complicated and the computations can be time-consuming. You can only understand these algorithms fully by implementing them, so that you can test the effect of varying data and parameters. Hence a successful completion of the course (and exam) requires that your implementation works and that you demonstrate a good understanding of the strengths and weaknesses of the models.
| No. | Date | Lecturer | Lecture | Exercise |
| 1 | 5 Sep | kvle |
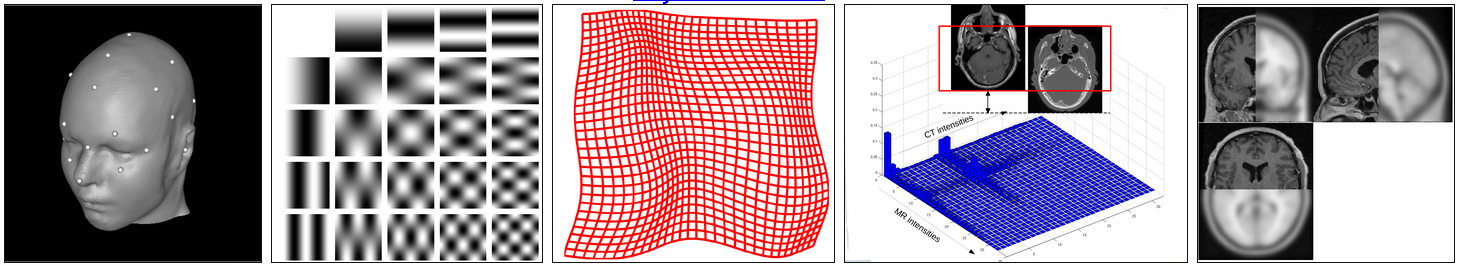
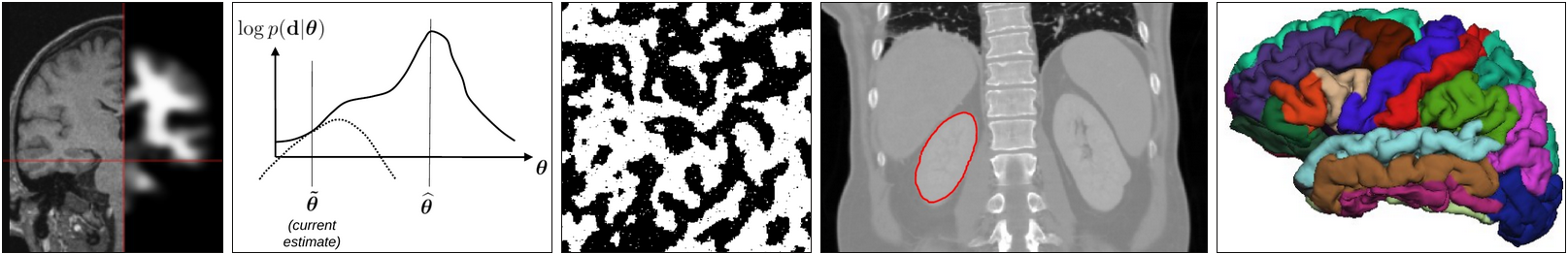
Introduction; Fitting of image functions (Course note pp. 1-14) Linear basis function models; maximum likelihood estimation and least squares; regularization; image smoothing; image interpolation |
MR bias field correction |
| 2 | 12 Sep | kvle | Landmark based registration (Course note pp. 15-24) | Landmark-based registration |
| 3 | 19 Sep | kvle |
Intensity based methods for registration (Course note pp. 25-32) Sum-of-squared differences; correlation; mutual information; principal axis transform |
Mutual information based registration |
| 4 | 26 Sep | kvle | Non-linear deformations (Course note pp. 32-46) | Non-linear intensity-based registration |
| 5 | 3 Oct | kvle | Surface-based segmentation (Course note pp. 55-55)
Active contour model; dynamic programming |
Non-linear intensity-based registration (cont.) |
| 6 | 10 Oct | kvle | Surface based registration (Course note pp. 51-55) | Surface-based registration |
| 7 | 24 Oct | Tron Darvann | EXCURSION: 3D laboratory. School of Dentistry | No exercise |
| 8 | 31 Oct | kvle |
Voxel-based segmentation I (Course note pp. 63-73) Gaussian mixture model; Markov random field priors; mean-field approximation |
Brain tumor segmentation |
| 9 | 7 Nov | kvle |
Voxel-based segmentation II (Course note pp. 73-85) Expectation-Maximization algorithm; MR bias field correction |
Expectation-Maximization algorithm |
| 10 | 14 Nov | Tim Dyrby | EXCURSION: Danish Research Center for Magnetic Resonance (DRCMR) | No exercise |
| 11 | 21 Nov | Martin Axelsen (Radiobotics) | Guest lecture | Expectation-Maximization algorithm (cont.) |
| 12 | 28 Nov | kvle |
Atlases (Course note pp. 93-102) Reference templates; group-wise registration; probabilistic atlases; label propagation |
Exercise catch-up |
| 13 | 5 Dec | kvle | Course wrap-up | No exercise |